Kozak Sequence
kiwifarms.net
- Joined
- Jul 3, 2025
Hi, all.
I'm making this thread because I am doing genetic engineering as a hobby and would like both: knowledgeable Kiwis' criticism of my own protocols I develop for my specific projects and for others to post their own projects' protocols or useful info in a format that is easy for others to derive relevant info from. I find a lot of information is either theoretical that is difficult to place in a protocol or is practical but is different from what I need and lacking relevant contextual information to make it useful. I'm still learning so if anyone else wants to share this flavor of autism with me it would be appreciated. Other science forums have been bled dry by Reddit and infested them with Reddit ideology. Redditors and their consequences have been a disaster for the human race, so here I post.
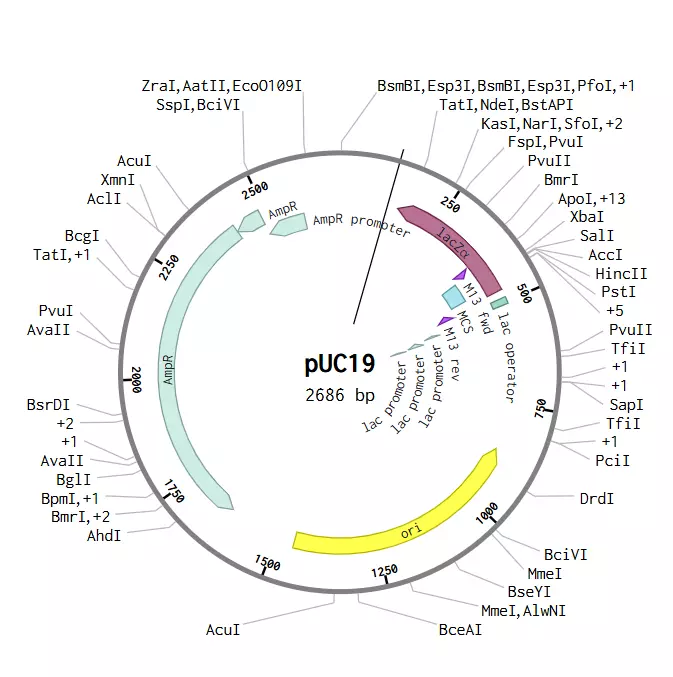
So this one is a simple transformation of DH5a E.coli to produce OxStayGold fluorescent protein(1) using PuC19 backbone. This strain isn't optimal for protein expression as DH5a is mostly used to archive plasmids but it's what I have in stock at the moment. for better expression you'd want one that can use the T7 promoter in many plasmids, like the BL21 strain(2).
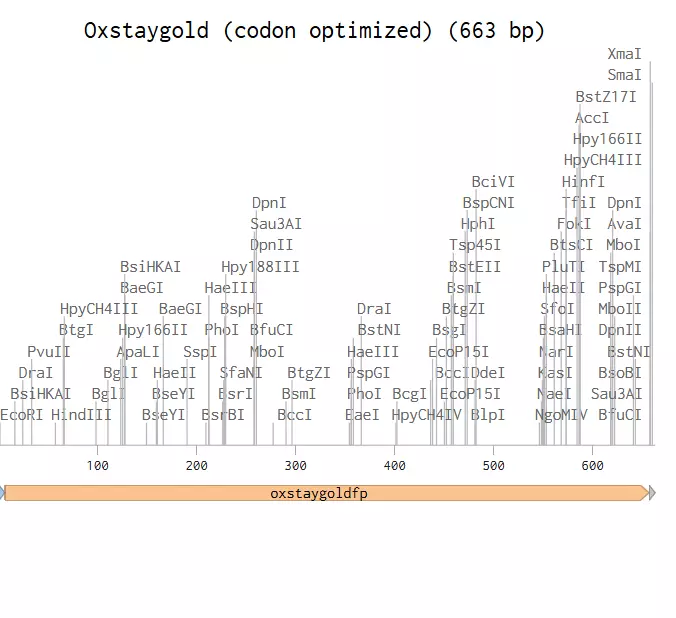

DNA Ordering and prep
You can order DNA from multiple companies. I got the following prep protocol from Twist Biosciences(3). The Thought Emporium has a video going over how to order it on Nebula which you can grab with a free trial(4).
- Upon receipt, briefly centrifuge tube or plate and resuspend the DNA in nuclease free Tris-EDTA (TE) buffer, pH 8.0 or 10 mM Tris-HCl, pH 8.0 to the desired concentration.
- We do not recommend resuspension in water.
- A concentration of at least 10 ng/ul is recommended for a stock solution, but optimal concentration will need to be determined based on your desired application.
- Prepare aliquots of the stock and separate working aliquots to limit the chances of contamination and reduce the number of freeze/thaw cycles.
- For long term storage, freeze DNA at -20° C for up to one year, or at -80° C for even longer storage.
E.coli broth and plating protocol:
Prepare LB broth liquid culture:
Materials
-Tryptone powder (5g)
-Yeast extract (2.5g)
-NaCl (5g)
-MiliQ water (500 ml)
Protocol:
1. Add the tryptone (5g), yeast extract (2.5g), and NaCl (5g) into a plastic beaker,
along with 400 ml of the MQ water. Use a magnetic spinner to dissolve.
2. Add 100 ml MQ water and further mix until dissolved.
3. Pour into a 500 ml Duran bottle and autoclave (15 psi, 121 124C, 15 mins)
Create antibiotic agar plates following same protocol except add 1.75G agar and add 100uG/ml ampicilin through a filter once it's below 55C. Keep in mind agar solidifies at 42C.
Comments: Though it says to use MiliQ water I'm fairly sure regular distilled water will work. Also make sure the syringe filter you use is hydrophilic. Hydrophobic ones will not filter many things and some are not clearly marked as such.
Competency protocol
Step 1: Inoculation and Growth of Cells (3-4 hours)
- Commence by inoculating a single colony of E. coli_ into 5-10 ml of LB broth.
- Incubate the culture at 37 °C with shaking until it reaches an optical density (OD600) of about 0.4–0.6. Should reach this in 2-3 hours.
Step 2: Chilling Cells on Ice (10-15 minutes)
- Transfer the culture to a sterile centrifuge tube.
- Place the tube on ice for 10-15 minutes to cool down the cells.
- transfer to centrifuge with thick gloves to keep temp low.
Step 3: Centrifugation and Resuspension of Cells (30 minutes)
- Centrifuge the culture at 4,000 x g for 10 minutes at 4 °C.
- Carefully decant and discard the supernatant.
- Gently resuspend the cell pellet in an equal volume of cold, sterile 100 mM calcium chloride (CaCl2) solution.
- Incubate on ice for 30 minutes, gently inverting the tube every 5-10 minutes.
Step 4: Repeating Centrifugation and Resuspension (30 minutes)
- Centrifuge the cells again at 4,000 x g for 10 minutes at 4 °C.
- Carefully remove the supernatant, leaving the cell pellet.
- Resuspend the pellet in a smaller volume (usually 100-200 μl) of cold 100 mM CaCl2 solution.
Step 5: Aliquoting and Freezing (15-20 minutes)
- Divide the competent cells into small aliquots (usually 20-100 μl) and store them in sterile microcentrifuge tubes at -80C if you are not immediately using them.
-Competent cells rapidly lose efficiency at -20C storage.
Comments: The most difficult part is keeping the centrifuge cold for 20 minutes. If you don't have a refrigerated centrifuge you can likely get away with putting yours in a cooler filled with ice, just make sure it's protected from meltwater. I also don't know why -80C is so fucking important but I'll take their word for it.
DNA preparation
After ordering from Twist, follow this protocol for both the backbone and the gene of interest. This uses specific restriction sites in PuC19 that I chose to use since they form sticky ends and have the same incubation temp and are both heat inactivated. These restriction enzyme protocols can be found at NEB(4).
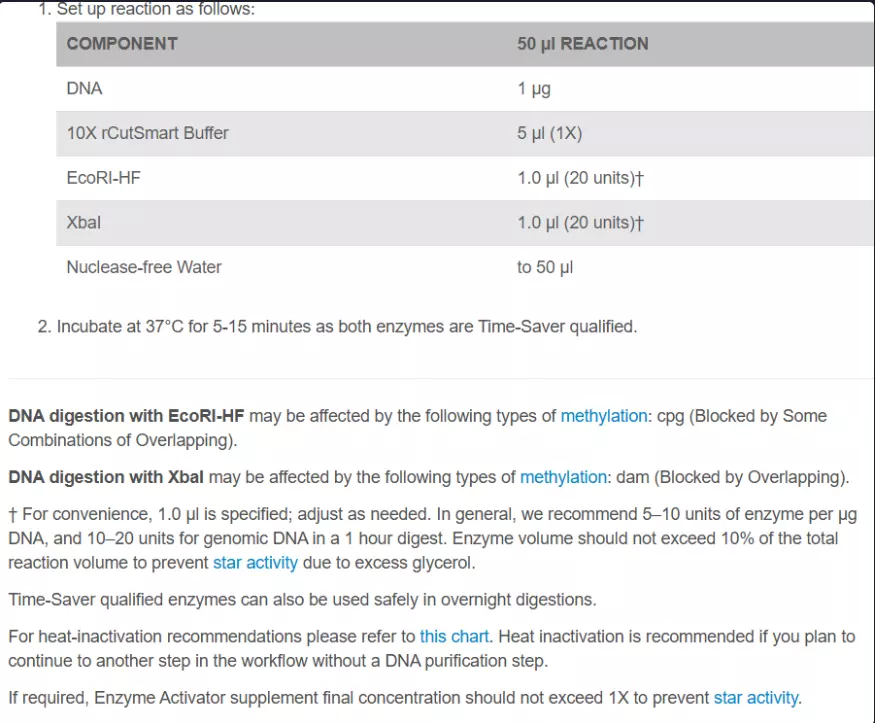
3. Heat inactivate at 65C for twenty minutes after incubation.
4. Cool to room temp.
CIP For just the backbone to prevent re-ligation of the cut out sequence. (Optional, most useful for blunt ends. Sticky ends have more difficulty in self-ligating)
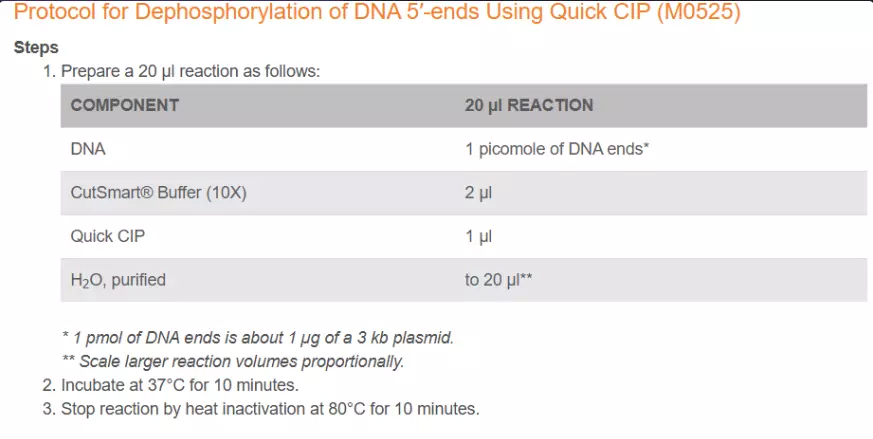

Ligate
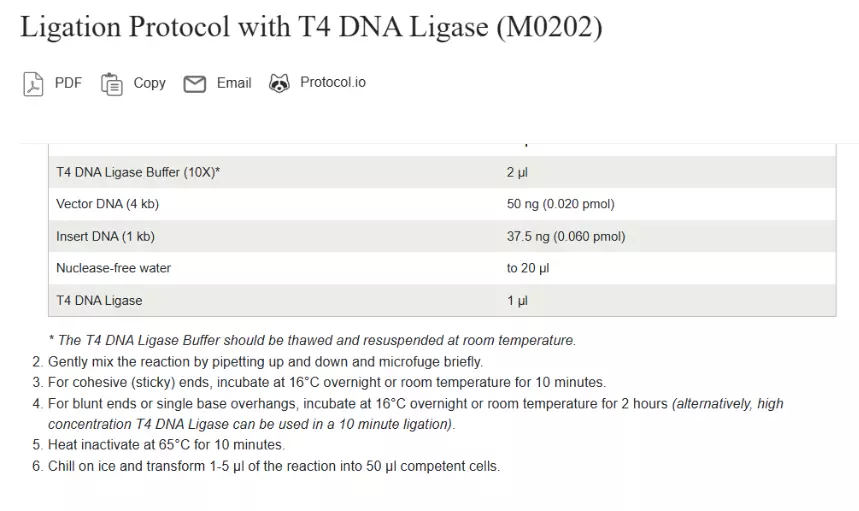
Transformation Protocol
Make sure laminar flow is available at the time of spreading (about 1.5-2 hr from the beginning of protocol).
1. Make sure the incubator is available for the entire duration of the experiment and is set for 37°C.
2. Keep the LB agar plates (with appropriate antibiotic) and media (or appropriate media) at room temperature or 37°C.
3. Water bath or heat block at 42°C.
Duration of the experiment
the experiment takes nearly 2 hours to complete the transformation, 12-16 hours after transformation (spreading)for the colonies to appear on the plate.
Procedure
1. Take out the competent cells from -80◦C and keep them on ice for 5-10 mins.
2. Add the ligation mix or plasmid to the competent cells and mix gently. Do not shake or vortex as it will inhibit transformation efficiency.
3. Incubate the reaction mix on ice for 10-15 minutes.
4. Give a heat shock to the cells by placing the reaction mix at 42°C for 30-90 seconds (water bath or Heat-block).
5. After the heat shock, transfer the cells onto the ice and add 500uL of warm LB
6. Place the tubes in the shaker (180 rpm) at 37°C for 1 hour.
7. After the incubation, give a brief spin at 4000 rpm for 2-3 minutes and decant the supernatant of 400 ul. Mix the pellet with the remaining supernatant (~100uL)
8. Split the mixture into 10uL and 90uL to spread onto different plates with appropriate antibiotics (or auxotrophic selection).
9. Incubate the plates overnight at 37°C.
10. After 12-16 hours of plating, colonies can be observed on the plates.
Variations of the protocol
Step 5: Adding 250 ul of media to the heat-shocked cells eliminates the spinning step, and one can directly spread the 10 % and 90 % volume on selective plates after 1-hour incubation.
From here you can add IPTG to trigger production of the protein.
This is what I put together for this specific project without getting into plasmid design, which if you want to know more you can look at the addgene blog(6) but there's a lot of small considerations that no one really talks about, like why the Multiple Cloning site of PuC19 is in the middle of the Lacza sequence, but its a standard plasmid so I assume its functional like that. This is partially why I'm posting here, because I don't have access to professors and all the books don't answer my questions. I hope this will be fruitful.
Citation:
1. https://www.fpbase.org/protein/oxstaygold/
2. https://www.embl.org/groups/protein...protein-expression/e-coli-expression-strains/
3. https://www.twistbioscience.com/faq/gene-synthesis?active=1016
4. https://nebcloner.neb.com/#!/redigest
5. https://nebula.tv/videos/thethoughtemporium-programming-dna-on-the-computer
6. https://info.addgene.org/plasmids-101-topic-page
I'm making this thread because I am doing genetic engineering as a hobby and would like both: knowledgeable Kiwis' criticism of my own protocols I develop for my specific projects and for others to post their own projects' protocols or useful info in a format that is easy for others to derive relevant info from. I find a lot of information is either theoretical that is difficult to place in a protocol or is practical but is different from what I need and lacking relevant contextual information to make it useful. I'm still learning so if anyone else wants to share this flavor of autism with me it would be appreciated. Other science forums have been bled dry by Reddit and infested them with Reddit ideology. Redditors and their consequences have been a disaster for the human race, so here I post.
So this one is a simple transformation of DH5a E.coli to produce OxStayGold fluorescent protein(1) using PuC19 backbone. This strain isn't optimal for protein expression as DH5a is mostly used to archive plasmids but it's what I have in stock at the moment. for better expression you'd want one that can use the T7 promoter in many plasmids, like the BL21 strain(2).
DNA Ordering and prep
You can order DNA from multiple companies. I got the following prep protocol from Twist Biosciences(3). The Thought Emporium has a video going over how to order it on Nebula which you can grab with a free trial(4).
- Upon receipt, briefly centrifuge tube or plate and resuspend the DNA in nuclease free Tris-EDTA (TE) buffer, pH 8.0 or 10 mM Tris-HCl, pH 8.0 to the desired concentration.
- We do not recommend resuspension in water.
- A concentration of at least 10 ng/ul is recommended for a stock solution, but optimal concentration will need to be determined based on your desired application.
- Prepare aliquots of the stock and separate working aliquots to limit the chances of contamination and reduce the number of freeze/thaw cycles.
- For long term storage, freeze DNA at -20° C for up to one year, or at -80° C for even longer storage.
E.coli broth and plating protocol:
Prepare LB broth liquid culture:
Materials
-Tryptone powder (5g)
-Yeast extract (2.5g)
-NaCl (5g)
-MiliQ water (500 ml)
Protocol:
1. Add the tryptone (5g), yeast extract (2.5g), and NaCl (5g) into a plastic beaker,
along with 400 ml of the MQ water. Use a magnetic spinner to dissolve.
2. Add 100 ml MQ water and further mix until dissolved.
3. Pour into a 500 ml Duran bottle and autoclave (15 psi, 121 124C, 15 mins)
Create antibiotic agar plates following same protocol except add 1.75G agar and add 100uG/ml ampicilin through a filter once it's below 55C. Keep in mind agar solidifies at 42C.
Comments: Though it says to use MiliQ water I'm fairly sure regular distilled water will work. Also make sure the syringe filter you use is hydrophilic. Hydrophobic ones will not filter many things and some are not clearly marked as such.
Competency protocol
Step 1: Inoculation and Growth of Cells (3-4 hours)
- Commence by inoculating a single colony of E. coli_ into 5-10 ml of LB broth.
- Incubate the culture at 37 °C with shaking until it reaches an optical density (OD600) of about 0.4–0.6. Should reach this in 2-3 hours.
Step 2: Chilling Cells on Ice (10-15 minutes)
- Transfer the culture to a sterile centrifuge tube.
- Place the tube on ice for 10-15 minutes to cool down the cells.
- transfer to centrifuge with thick gloves to keep temp low.
Step 3: Centrifugation and Resuspension of Cells (30 minutes)
- Centrifuge the culture at 4,000 x g for 10 minutes at 4 °C.
- Carefully decant and discard the supernatant.
- Gently resuspend the cell pellet in an equal volume of cold, sterile 100 mM calcium chloride (CaCl2) solution.
- Incubate on ice for 30 minutes, gently inverting the tube every 5-10 minutes.
Step 4: Repeating Centrifugation and Resuspension (30 minutes)
- Centrifuge the cells again at 4,000 x g for 10 minutes at 4 °C.
- Carefully remove the supernatant, leaving the cell pellet.
- Resuspend the pellet in a smaller volume (usually 100-200 μl) of cold 100 mM CaCl2 solution.
Step 5: Aliquoting and Freezing (15-20 minutes)
- Divide the competent cells into small aliquots (usually 20-100 μl) and store them in sterile microcentrifuge tubes at -80C if you are not immediately using them.
-Competent cells rapidly lose efficiency at -20C storage.
Comments: The most difficult part is keeping the centrifuge cold for 20 minutes. If you don't have a refrigerated centrifuge you can likely get away with putting yours in a cooler filled with ice, just make sure it's protected from meltwater. I also don't know why -80C is so fucking important but I'll take their word for it.
DNA preparation
After ordering from Twist, follow this protocol for both the backbone and the gene of interest. This uses specific restriction sites in PuC19 that I chose to use since they form sticky ends and have the same incubation temp and are both heat inactivated. These restriction enzyme protocols can be found at NEB(4).
3. Heat inactivate at 65C for twenty minutes after incubation.
4. Cool to room temp.
CIP For just the backbone to prevent re-ligation of the cut out sequence. (Optional, most useful for blunt ends. Sticky ends have more difficulty in self-ligating)
Ligate
Transformation Protocol
Make sure laminar flow is available at the time of spreading (about 1.5-2 hr from the beginning of protocol).
1. Make sure the incubator is available for the entire duration of the experiment and is set for 37°C.
2. Keep the LB agar plates (with appropriate antibiotic) and media (or appropriate media) at room temperature or 37°C.
3. Water bath or heat block at 42°C.
Duration of the experiment
the experiment takes nearly 2 hours to complete the transformation, 12-16 hours after transformation (spreading)for the colonies to appear on the plate.
Procedure
1. Take out the competent cells from -80◦C and keep them on ice for 5-10 mins.
2. Add the ligation mix or plasmid to the competent cells and mix gently. Do not shake or vortex as it will inhibit transformation efficiency.
3. Incubate the reaction mix on ice for 10-15 minutes.
4. Give a heat shock to the cells by placing the reaction mix at 42°C for 30-90 seconds (water bath or Heat-block).
5. After the heat shock, transfer the cells onto the ice and add 500uL of warm LB
6. Place the tubes in the shaker (180 rpm) at 37°C for 1 hour.
7. After the incubation, give a brief spin at 4000 rpm for 2-3 minutes and decant the supernatant of 400 ul. Mix the pellet with the remaining supernatant (~100uL)
8. Split the mixture into 10uL and 90uL to spread onto different plates with appropriate antibiotics (or auxotrophic selection).
9. Incubate the plates overnight at 37°C.
10. After 12-16 hours of plating, colonies can be observed on the plates.
Variations of the protocol
Step 5: Adding 250 ul of media to the heat-shocked cells eliminates the spinning step, and one can directly spread the 10 % and 90 % volume on selective plates after 1-hour incubation.
From here you can add IPTG to trigger production of the protein.
This is what I put together for this specific project without getting into plasmid design, which if you want to know more you can look at the addgene blog(6) but there's a lot of small considerations that no one really talks about, like why the Multiple Cloning site of PuC19 is in the middle of the Lacza sequence, but its a standard plasmid so I assume its functional like that. This is partially why I'm posting here, because I don't have access to professors and all the books don't answer my questions. I hope this will be fruitful.
Citation:
1. https://www.fpbase.org/protein/oxstaygold/
2. https://www.embl.org/groups/protein...protein-expression/e-coli-expression-strains/
3. https://www.twistbioscience.com/faq/gene-synthesis?active=1016
4. https://nebcloner.neb.com/#!/redigest
5. https://nebula.tv/videos/thethoughtemporium-programming-dna-on-the-computer
6. https://info.addgene.org/plasmids-101-topic-page
